Overview
In a post-genomic era, the importance of translational control in gene expression has been increasingly appreciated. Dysregulation of mRNA translation has been implicated in human diseases such as cancer, diabetes, and neurodegenerative disorders. Using biochemical, genetic, and genomic approaches, we are cracking the mystery of this fundamental cellular process.
Research Projects
 |
|
Translational Reprogramming in Gene Expression |
| |
|
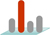
Enabling swift regulation in gene expression, translaional reprogramming can be quantitative (all-or-none vs. graded ), qualitative (enabling a single mRNA to produce several different proteins), or selective (activating subsets of mRNAs for translation). Translational reprogramming primarily involves in proper start codon selection, which plays a crucial role in cell growth, differentiation, stress response, as well as organismal development.
We developed genome-wide approaches for profiling of initiating ribosomes, which enables quantitative mapping of translational initiation sites across the transcriptome. Using eukaryotic cells and mouse models, we are actively investigating alternative pathways controlling ribosome loading, scanning, and start codon selection.
- Lee et al. PNAS 2012; 109(37):E2424-32.
- Gao et al. Nat Methods 2015; 12(2):147-53
|
 |
| |
 |
|
mRNA Epigenetics in Cellular Physiology |
| |
|
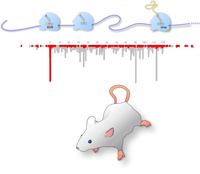
Much like the epigenetic modification of DNA and histones, mRNA is also subject to extensive modification. The dynamic RNA methylation heralds the arrival of RNA “epigenetics”. The most abundant mRNA modification is N6-methyladenosine (m6A), whose regulation and physiological significance are only beginning to be unfurled.
Using m6A-seq, we discovered dynamic regulation of mRNA methylation in response to cellular stress. By focusing on the role of m6A modification in alternative modes of mRNA translation, we are currently investigating the broad effects of mRNA methylation in cellular physiology, such as cell growth, development, as well as cancer.
- Zhou et al. Nature 2015; 526(7574):591-4
- Meyer et al. Cell 2015; 163(4):999-1010
|
 |
| |
 |
|
Stress Response in Protein Homeostasis |
| |
|
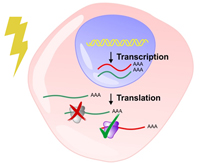
Cells rely on swift and selective regulation of gene expression to protect cells from adverse conditions. With translational control, global protein synthesis could by quickly shutdown, whereas a subset of mRNA undergoes selective translation to produce proteins essential for cell survival. Mechanistic details of translational regulation in response to stress remains incompletely understood.
Using heat shock and nutrient starvation as stress models, we are investigating functional alterations of ribosomes and initiation factors. We aim to uncover novel modes of translational regulation in response to stress.
- Liu et al. Mol Cell 2013; 49(3):453-463
- Zhang et al. Nat Struct Mol Biol 2015; 22(5):404-10
|
 |
| |
| |
|
|
| |
|
. |
| |
|
|
|



